On this page we'll build a neuron with specific connection geometry. On the next page we'll build a network with thousands of these neurons, and create some data with it. Before beginning though, I'd like to show you one quick thing, it's in the way of Annie being smart about networks. As already mentioned, you can spend the time up front to create a beautiful hierarchical network, with neurons inside layers of cells inside well defined nuclei. But reality doesn't always work out that way - sometimes you're working late at night and suddenly you have a Eureka! moment, and then you want to immediately save whatever you have - sometimes you don't even know what you have, you just need to save it so you can figure out later what you did to make it work. With Annie, you can just keep creating cells and neurons till you find something interesting, you don't have to organize them into networks and nuclei up front, you can just leave them dangling - but when you get to that Eureka! moment and you need to save your network, push the "Build Network" button and Annie will automatically generate a network for you. She'll move all the necessary pieces out of your client lists and into a system generated network. This function is so important, we'll show it here, up front. Here's how it works. Let's say we're just creating experimental cells and neurons. We create a cell, this way:
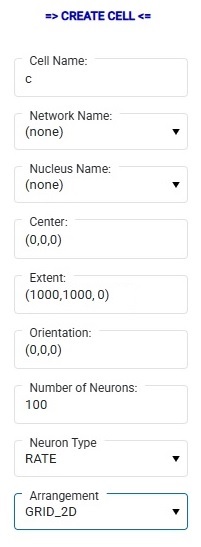
When we do this, the cell is "dangling", it isn't a part of any network because we haven't selected a network, the network drop-down is empty. But now we can populate this empty cell with neurons, and we can use them immediately in a simulation by pushing the "Build Network" button. Annie will then generate a network for us. Here's how that looks:
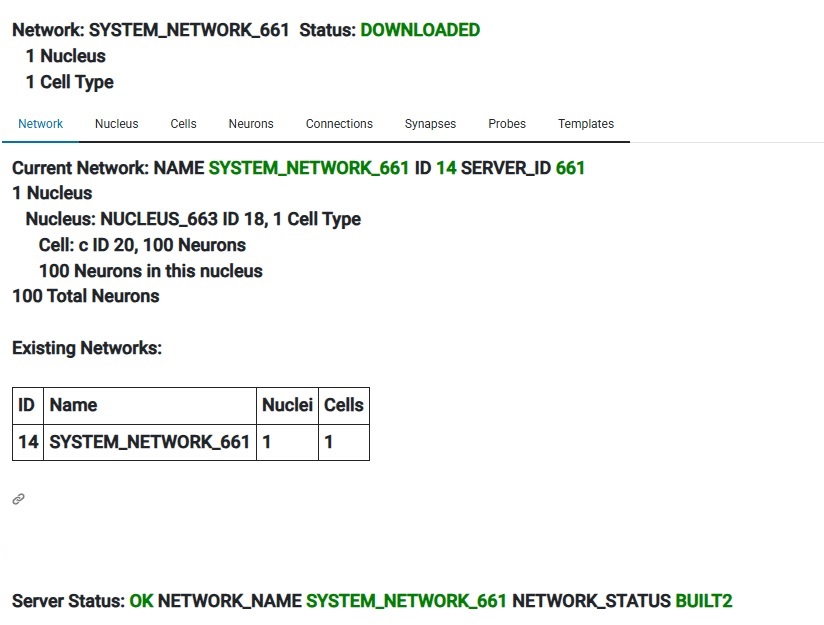
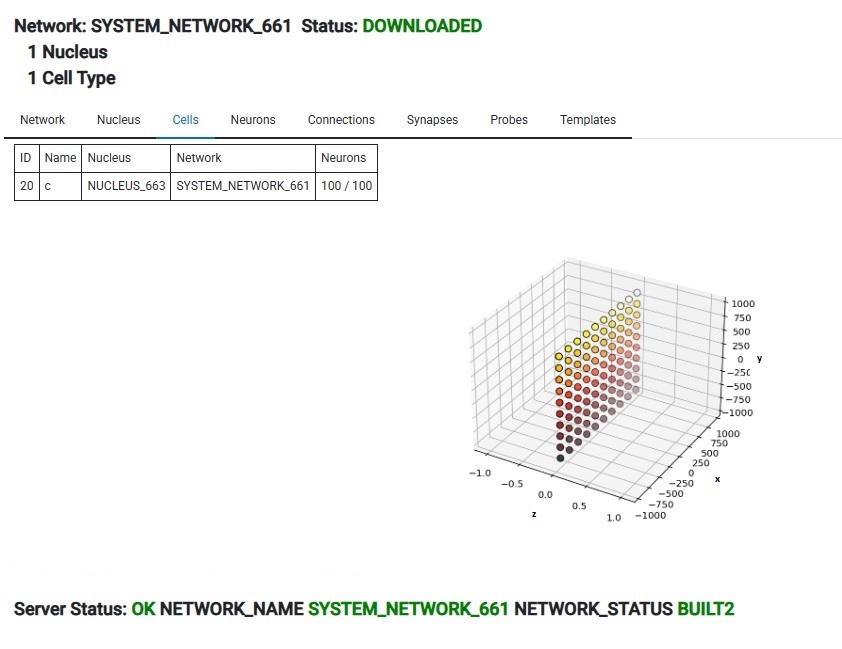
Annie has created a network and a nucleus for us, populated our cell with neurons, and moved the cell (called "c") and the designated 100 neurons into both the network and the nucleus. The network is called SYSTEM_NETWORK_661 and we can now rename it to anything we wish. When building a network this way, Annie will take "all" the dangling objects and include them in the new network, and if we don't want this, we can simply remove any cells we don't want. Or, we can do this entire process manually one cell at a time, we can create our own network and move cells into it and then build the network. Usually Annie can do it faster than we can. This feature appears in many workflows, perhaps the most important of which is the "emergency save" upon discovery or upon error. But now that we have a network, we can "Create Simulation" and then "Add Network To Simulation" and then push the Start button. That's how easy it is to run a simulation. Takes less than a minute to build a working simulation from scratch. (Annie's testers can do it in about 15 seconds, they're very good). Of course in this case nothing interesting will happen, because we haven't defined any connections. But you can easily see that this workflow can save plenty of time compared to trying to do the same thing in Brian2. Nest is a little friendlier, and so is Nengo, but Annie leaves the others far behind when it comes to the subsequent geometry, and in some cases even the computations. For those who care about geometry, Annie is an essential part of the toolset.
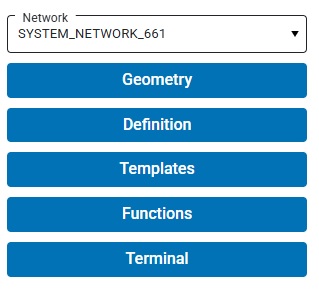
Annie maintains the concept of a "current network". The current network can be selected using the Network drop-down in the left sidebar. After a build, the built network appears in the dropdown. You can just keep creating networks and building them, you can have dozens of networks built and available to choose from. This is helpful when you're trying to compare the behavior of similar but slightly different networks.
Here's our neurons, that were built from the CELL we created. There's 100 of them, and Annie has positioned them on a 2-dimensional grid like we told her to. We can switch over to the Neurons tab and look at them in report form, or see the graphical layout in the Cells tab, where we can zoom, pan, rotate, and pick individual neurons for examination.
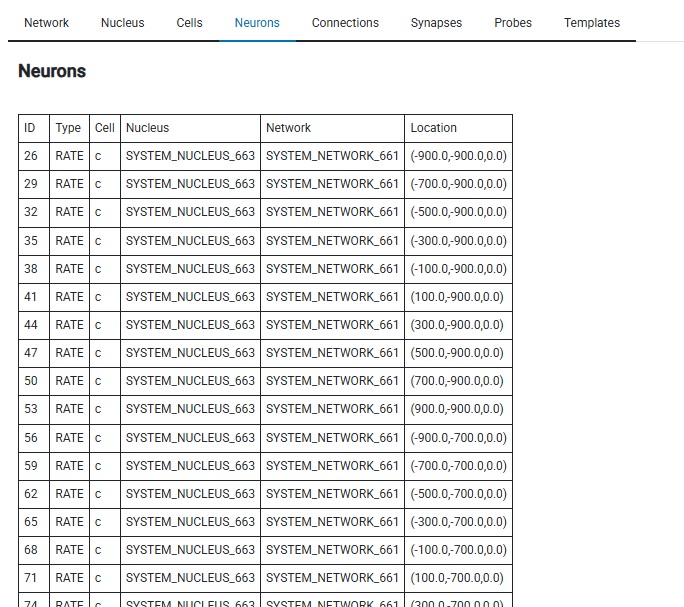

All fine and good. But... neurons as spheres? Puh-lease! That's so... 1940's! Thanks for being patient, I had to show you the simple interactive dashboard first, to provide context for what follows. Inside Annie's simple dashboard are some very powerful tools. For one thing, each tab is a VTK visualizer, what you see depends on what you're doing. The "Neurons" tab can show you a list of the neurons that belong to a cell group (and their locations, membrane properties, connections, and so on - in tabular form), but it can also show you a picture of a neuron, and let you zoom in on a dendrite all the way down to the level of individual ion channels. (No kidding!) We have to look at solid body modeling before showing computations on meshes. The mesh is the geometry, and there are numbers attached to the cells and vertices. The numbers could be scalars like membrane potential, or vectors like ion currents. To make the leap from solid bodies to meshes, we can look at some neural mesh geometry.
So okay, let's have some fun. And we'll be real and productive at the same time, by introducing the concept of meshes, and showing how Annie uses them to build geometry. In this exercise, we're going to create a neuron. Specifically, we're interested in the branching geometry of its axon. Let's say, this is a thalamocortical relay neuron, from the LGN to visual cortex V1. Its axon leaves the LGN in a bundle of other axons, arranged in a topographic manner, and eventually it reaches its target in the cerebral cortex, where it branches in a tuft in layer 4cβ. So our axon wants to travel through the optic radiation to a specific target location, and connect to a specific layer (and specific cells) within it. On the next page we'll run a few thousand of these neurons in a simulation. First though, let's build the prototype neuron.
The idea with Annie is that you can specify your geometry at any level of detail you wish. (You can also use a light theme or a dark theme and customize your user interface - the screen shots above show Panel's light theme, those below show VTK's dark theme. The network definition language is designed so that it can go all the way from the network container down to the individual positioning of ion channels in a membrane. Much of the time the extreme detail is not needed, however sometimes it is. VTK is a very powerful visualization tool, especially when combined with a mesh engine. For example, membrane channel positioning can be determined in several ways. The hard way is, you can actually detail each individual channel in the network definition file. An easier way is, you can tell Annie to lay out the channels for you according to a MAP. An even easier way is, you can read in a mesh and position the channels along the vertices. Here is an example of the power of VTK - and this is just an example (there are many impressive examples on the VTK home page). First, we can envision a small section of axon, with a cytoskeletal framework. We can specify this as a vector with an orientation and a diameter, and Annie (or Blender or VTK) will create a cylindrical mesh for us.
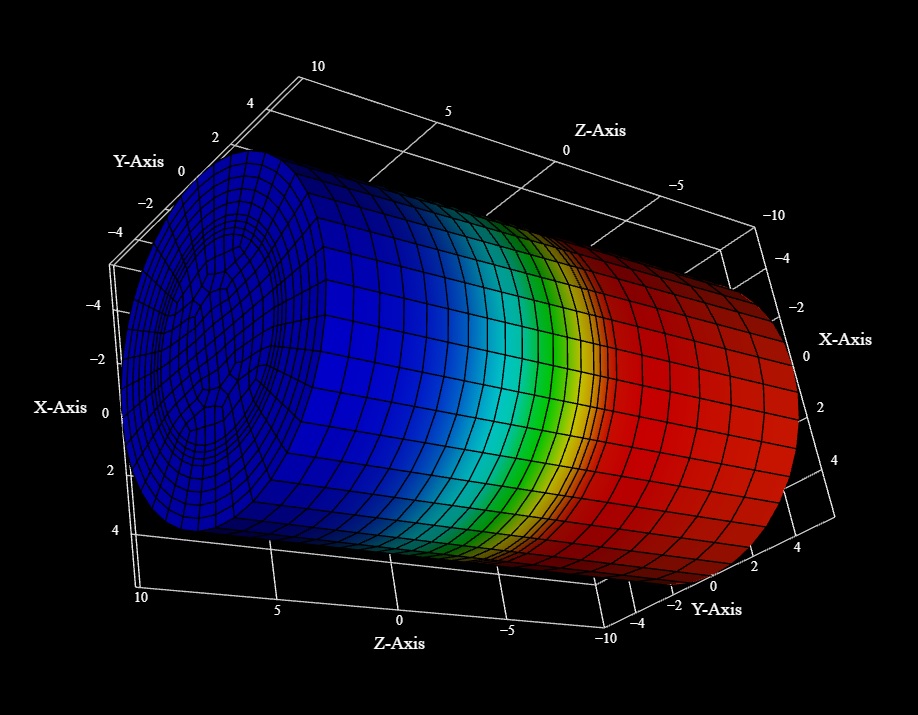
Upon import, Annie can position ion channels at the vertices, and generate a reasonable facsimile of a cytoskeleton by understanding which part of the mesh is the inside. These view are fully interactive, and you can then pick and select individual channels (vertices) simply by clicking on them. VTK is just showing us the mesh (it can be displayed as a wire frame or shaded as a solid object), there's nothing much to it. But you can see how the interaction between the mesh and the network can be a powerful tool. To determine where the ion channels are, we can simply set the mesh resolution. Easy. 60 seconds. 45 seconds to create the mesh, and another 15 seconds to import it and visualize it. The channel properties are defined by the components of the NEURON, which inherit from the components of the CELL. You can build a prototype at the CELL level, and transmit it down to the neurons with variance, and this way instead of the regular geometry shown here, Annie will position the channels "around" the vertices and create a realistic membrane. However these images suffice to demonstrate the concept and the workflow.
To see what the mesh actually looks like, we can switch over to wire frame mode by simply pushing the 'w' key on the keyboard. A careful examination of the wire frame reveals an outer membrane with 5 layers, where transmembrane proteins can be positioned on one side of the bilayer or the other. The green structure could be a piece of endoplasmic reticulum in a dendritic spine, as it seems to be defining a structural channel along the -Z axis. These structures can be used for any purpose whatsoever, they merely define geometric locations, which are points of attachment for data. Every vertex in the mesh has data attached to it, it could be a membrane potential, or an RGB value for display, or any other useful information. And significantly, the data can include equations. Annie has an enormous list of built-in functions that you can attach to any object, in the membrane or otherwise. These equations can include ion channel kinetics, channel diffusion in the membrane, influences from nearby currents and voltages, and even long-distance interactions defined as "coupling" (which could be fields or any other thing but generally are related to covariances with remote states). The simulator simply runs down the list of network equations and calculates each in turn, it doesn't care what's attached to what except insofar as it can access the variables.
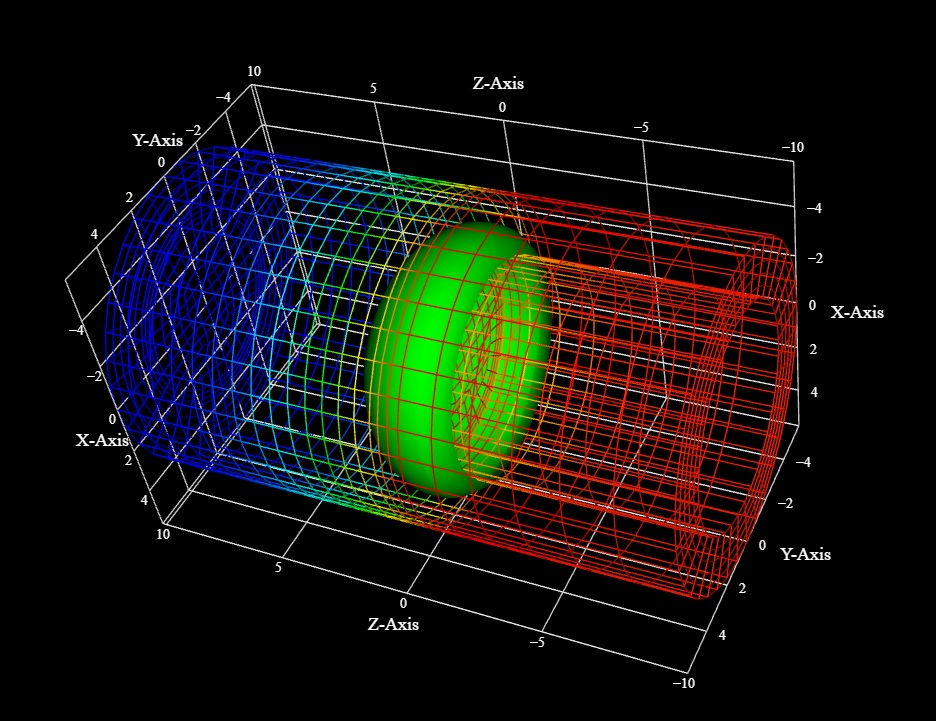
VTK is a fast and powerful visualization tool. When rendering locally, there's a lot of data being transmitted across the network (geometry is downloaded from the server into the workstation). Typically a large network (100,000 neurons) takes a few seconds in client-server mode, and if you're working on detailed mesh geometry there may be some further transfer. Annie can, however, operate entirely in stand-alone mode, without a server. All the same capabilities are in place, however large networks will eat up workstation memory in this mode, especially when using the mesh tools. The stand-alone mode is designed for casual use, when you don't need the full power of the server. For example you could run ordinary neuron-level simulations without using the mesh engine. There is also a "mirror" mode, which is enormously helpful for debugging. In this mode, the workstation will replicate everything the server does, which reduces the clutter on the wire and also uncouples the local network. These modes are helpful when building networks bottom-up, you can work on membrane geometry till you're happy, and then when you're ready you can build neurons and a network.
Let's look at how we get from "neurons as spheres", to "neurons as meshes". To build a neuron, we can invoke Annie's geometric capabilites. In this case we'd like to build an axon, and the same principles apply to building dendrites. The difference between axons and dendrites is there's usually only one axon, whereas there can be many dendrites branching directly off the cell body. Generally we're interested in the shape of the "branching tree" associated with these processes. There are many ways to ask Annie to build a tree for us. The general algorithm is that the axon travels forward a certain distance, then branches, and the branching is characteristic. From the standpoint of fractal geometry this is a straightforward procedural algorithm and all we have to do is specify the diameters of the branches. We can provide Annie with three arrays: one that tells her when to branch, another than tells her how, and a third that tells her how much variance to apply to the branchpoints. This results in very realistic branching geometry. With Annie, segments of membrane ("compartments") are defined as vectors, with data attached to them. To generate a branching axon, you can pass Annie a VECTOR, that serves are the root of the tree, and a set of of branching instructions. Vectors can be relative or absolute. In relative mode, they are specified the same way POINTs are (the origin is stipulated to be 0,0,0). In absolute mode, vectors need a beginning point and an end point. VECTORs are different from points, in that they can have data attached to them. In this case, the vector is associated with the diameter of a cylinder. To define a branching tree for an axon, we first tell Annie where to start. Then we pass in a branching spec, that defines the branch points, and what to do at each point. For example here is an AXON as it would be specified in a network definition file.
CELL XYZ
...
NEURON_SIZE (5,5,5)
...
AXON (5,5,0) CYLINDER * this tell Annie where on the neuron to position the axon
BRANCH [ [(0,0,4000)], [(200,10,0),(200,-10,0)], [(100,10,0),(100,-10,0),(100,-10,10)] ]
DIAMETERS [ 100.0, 80.0, 60.0 ]
Here we're telling Annie to position the axon at a particular location on the neuron cell body, and we can specify additional parameters like ORIENTATION. Without modifiers, the axon will travel straight out from the neuron for a distance of 4000 units along the Z axis (the first item in the BRANCH array), where it will branch. It will split into two branches, because there are two items in the second element of the BRANCH array. After it splits, the second branch will be very short, it will only travel for 200 additional units before it branches again, into 3 pieces. And, the DIAMETERS array specifies the diameter of each segment. So here, we have "hard coded" a branching specification. There are easier ways to do it, for instance a TREE directive or a FRACTAL directive, but this suffices for purposes of illustration, and ultimately all of the higher level directives are converted to detailed branching specifications like the above. When built this way, the branches naturally dovetail with functional compartmentation. Each cylindrical segment can be computed individually, and transmission through the branchpoints can be controlled algorithmically.
That's the "manual" way to do it, we can actually pass Annie the arrays and ask her to build accordingly - on the other hand, if we already have all the specs ahead of time then we hardly need Annie, we can just build our own neurons. One of the neat things Annie can do is move axons around. For instance in the earlier example on the previous page, some "random paths" were shown that ended up traveling directly through neurons and such. We can prevent this by collecting the axons into a BUNDLE and specifying the bundle geometry. We can also simply use an AVOID_COLLISIONS directive when building the connections, which works better in close situations when bundles don't apply. In the context of an optic radiation, we'll probably want to define a bundle, that way we can do slick things with the visualizations, like color-code the topography and so on. Bundles are defined by loops through which the axons have to travel, they are geometric constraints.
The problem with visualizing long axons is they're tiny! The neuron may only be a few microns wide, and if we attach an axon that's a micron wide and ask it to travel 10 cm, it will be hard to visualize, because it'll look like a tiny little thread in a sea of ... whatever else is there. Here's an example, it's very hard to see the axon in a large coordinate space but when we zoom in everything's fine.
 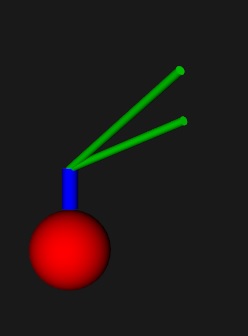
To alleviate this issue you can click on a neuron and identify its processes, but they can still be small and hard to see. So Annie will identify the endpoints for you, making it easier to zoom in to the location of your choice. Once you've identified the endpoints you can look at the axon as a big fat tube, making it very easy to pick and select. The axons are actually very fine meshes, as you can see here:

The mesh resolves down to about 10 Angstroms, which is sufficient to visualize ion channels. The resolution is actually arbitary, but at some point becomes limited by the representation of a floating point number inside the computer. In the supercomputer world we have 128-bit IEEE floating point math that's good down to 35 decimal digits, while with ordinary 64-bit floats like Python has we can do 15 or 16 digits. The real resolution is about half of that, because when you multiply two small numbers together the result should remain non-zero. It's hard to pick vertices by looking at an axon, you have to zoom in repeatedly to small patches of membrane before you can see the underlying mesh. Sometimes it's easier to choose members from an object list, especially when you can name objects so you can find them later. That's also one of the nice things about bundles, you can pick a bundle and suddenly you can see the underlying geometry (although even in bundles, the fiber pathways can be exceedingly small, but at least you have something to click on and make bigger). Annie can help by colorizing your axons and dendrites. Anyway, arrays are a difficult way to specify branching geometry. The good news is, Annie has libraries of canned neurons! You can choose a shape from the library, and endow it with the channel structure of your choice. Or start with a shape from the library, and modify it as you wish. Cell bodies can be stellate, pyramidal, or fusiform, and there are basket cells, chandelier cells, all manner of interesting geometries, built from simple branching primitives. Choose a spherical or pyramidal cell body, specify your branching lengths and diameters, and you're done. You can do it interactively, from a definition file, from a picture, from other simulators, even from CAD/CAM and 3-D printing software.
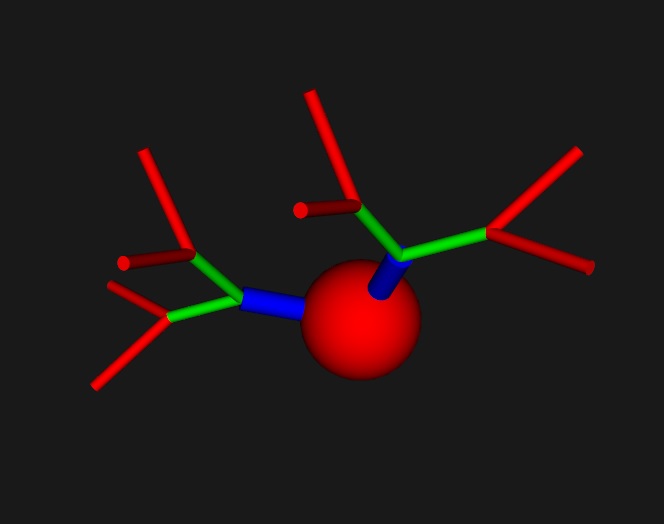 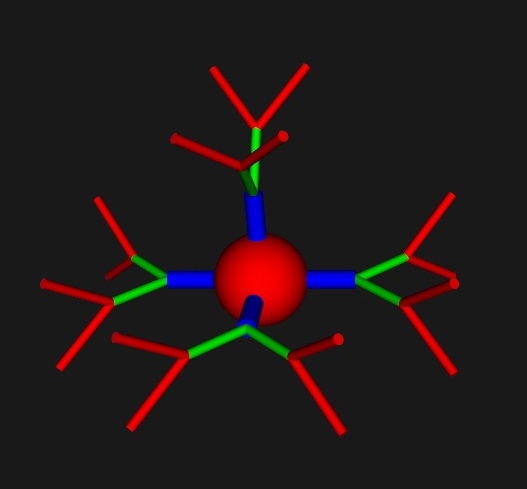
These neurons may look primitive, and in fact they are. (I have to show you the simple stuff first). The magic happens when we work with the meshes a bit. Annie knows about the NAVis tool, and can import its meshes. Janelia has some wonderful Drosophila meshes, Allen has lots of mouse brains, and you can trace neurons into Blender and mesh them up that way. Actually though, the simple geometric shapes are very useful. (And they come without the zillions of dependencies that NAVis requires). You can take any of the primitive shapes, shorten or lengthen the processes, and send them off in different directions. In a way, they're exactly like VTK pipelines, the primitives are the Sources, and the modifiers are the Filters. This is an example of a NAVis neuron:
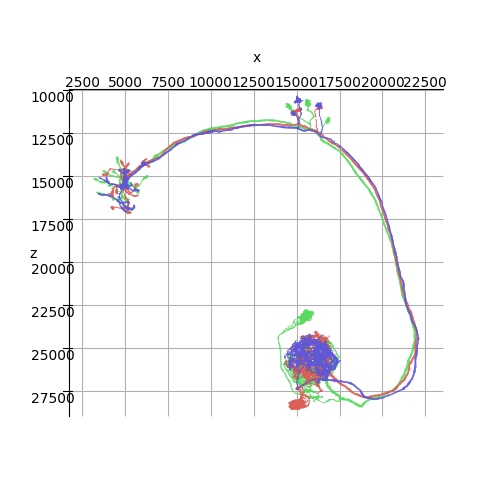
It's nice as far as it goes, but Annie can do the same thing with a mesh that allows you to zoom all the way in to a patch of membrane and visualize the ion channels. And of course VTK is a whole lot prettier than matplotlib. Obviously, detailing a large network at this level is expensive in terms of computer memory, however there is no limit to the size of a mesh and if you have the gB why not use them! You can do a lot with 32 gB, which most laptops have these days. I've seen people use Maya on laptops with a 14" screen, I can't do it myself (especially while walking around) but it's certainly possible. This is why we need VTK, Annie has been working in text mode for 20+ years and never had a dashboard until now. People were satisfied with PNG's and CSV files. (But I'm not - unfortunately the interactive part of VTK is still crashing once in a while, and I'm working very hard on a daily basis to ensure that Annie's fans have a delightful user experience when they test the dashboard for the first time. So far I can just give you a glimpse of what Annie is about, and I don't want to show you a gazillion lengthy text files so I'll just keep updating this web site as things get done). Anyway, meshes are the answer to a lot of annoying issues in the neural modeling space, and the specific reason we like meshes, even more than geometry - is computation. These are computational meshes, every vertex has data attached to it. Compartments are kind of a primitive way of dealing with axons and dendrites, more likely they're like one of Ilya Prigogine's experiments in nonlinear thermodynamics, where every point is an oscillator and one needs to model the coupling on a lattice before one sees the covariances that contribute to the spatial patterns.
Sometimes meshing a picture is faster than building a branching tree from scratch. Another possibility is, that rather than giving Annie arrays, we can ask her to build them for us. The heuristics of fiber pathways are actually pretty simple, with a few easy rules we can put together realistic geometry. One of the important concepts that other simulators fail to address is the neuropil. Neuropils are ubuiqitous in brains, they're basically layers of synapses, they're well defined geometric structures where axons and dendrites from various neurons comingle and interconnect. For example in the retina there are inner and outer plexiform layers. The inner plexiform layer is probably a classic neuropil, it connects 12 kinds of bipolar cells, 60+ kinds of amacrine cells, and 20+ kinds of ganglion cells. The connectivity within the neuropil is determined chemically, on the basis of markers expressed by the cell types. It's not perfect, it's probabilistic - synapses connect "with a probability", and sometimes the connection probability doesn't go entirely to zero (in real brains we frequently see oddball synapses that seem to go outside the mainstream connection patterns).
The easiest way of all, of defining axon geometry, is simply to draw it on the screen. You can use Annie, or any mesh tool. All you need is a prototype, and you can get that from a picture, you can easily trace a Golgi stained image into Blender and import the mesh directly into Annie. When Annie imports your axon, you can tell her how to treat your vertices, she can put a branch point at every vertex, or she can use groupings in the OBJ file. These are advanced topics, but I'm tying to give you a flavor for how powerful Annie is. You can create axons many different ways, define them as text, import them from Blender, use functions to define their trajectories and constrain them into bundles, or simply draw them on the screen. With the model axon as given, the results are not very exciting, but they suffice to illustrate the principle. Here's our axon with the size of the first segment reduced so it can be visualized. We created this from scratch just now, we didn't draw it from the library. In the CELL spec we told Annie about the NEURON_SIZE and in the branching spec we told her where the AXON attaches and how to generate the branches. It wasn't hard, right? Three lines in a definition file? "This is where the axon starts, this is how it branches, and these are the diameters". Easy.
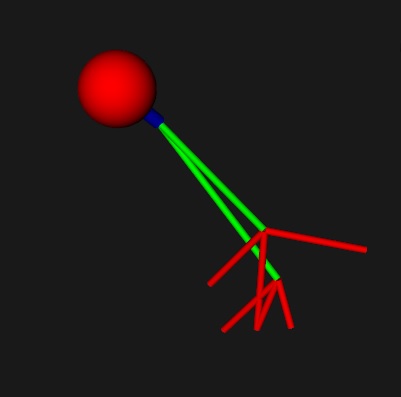
We defined this axon at the level of the CELL, so it can now be transmitted to the individual neurons, and if we add a line like this to the branching specification:
VARIANCE [ (0,0,0), [(2,2,2),(2,2,2)], [(1,1,1),(1,1,1),(1,1,1)] ]
we can get a nice array of similar but slightly different neurons. This geometry is used in the calculation of synaptic positioning and neuropil structure. Here we have built an axon, but we've said nothing about how it's connected. That part belongs to the CONNECTION specification, and we'll get into that in a moment (finding nearest neighbors in a neuropil is no different than mapping them from layer to layer, it can be done the same way, including a mapping function or a MAP). The Lego-style building-block approach to neural geometry is actually very powerful. One can generate branching geometry independently of neural positioning, attach branching styles to the cells of our choice, and so on. ("Hey Annie, go put this axon on a neuron, then give me 100 of them with a slight variance laid out in a 2-d grid"). Annie will get it done faster than you can blink your eyes.
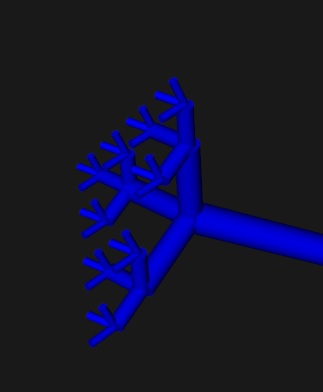

Computationally, this neuron will behave like any other neuron till we tell Annie that it's special. Visualizing the neuron's geometry suggests some interesting possibilites, doesn't it? For one thing, the cylindrical visualization reminds us of myelinated motoneurons, and we can in fact ask Annie to generate some Nodes of Ranvier for us by wrapping some Schwann cells around cylindrical segments of the axon. We can then define in detail the computational relationship between the axon segments and their associated glia. I don't want to get into the computational piece just yet, because it's complicated and it takes multiple web pages to explain (it's even hard to do justice to Annie's geometric capabilities on a web page, but I'm trying). Yes, Annie can do all the things Brian2 can do, but the equations are in a much cleaner and user-friendly form, and you can use commonly available tools to translate your own equations into terms Annie can understand. That's what the "Functions" button is for, below the Network selector in the above pic.
The model neurons on this page are very simple examples, in real life we'd wish to elaborate these considerably. For instance here are some real neurons, the retina is on the left (you can see the beautifully layered neuropil), and a cortical chandelier cell is on the right.
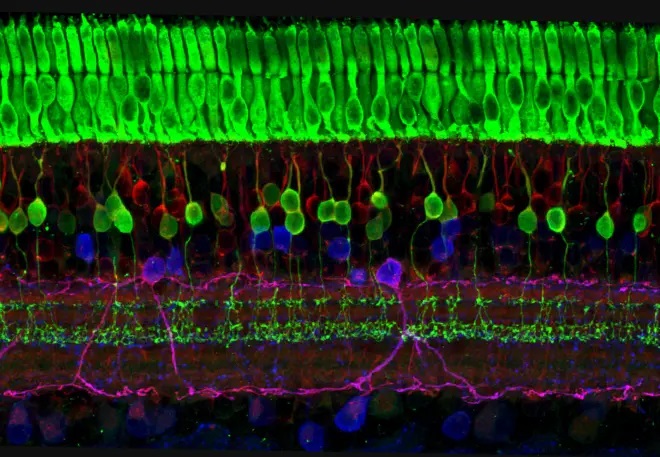 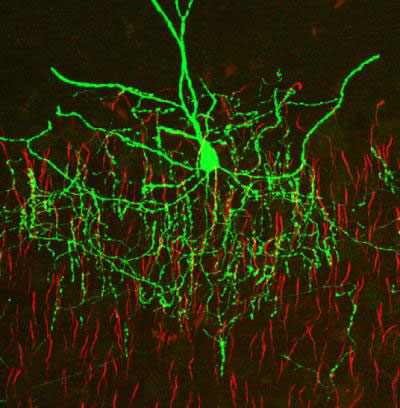
It would take all year to generate such a neuron manually, and the good news is there are other easier and faster ways to do it. Here's one of them, you can simply trace the picture. You can import your picture into Blender, trace the outlines, and Annie will fix the mesh and build a neuron.
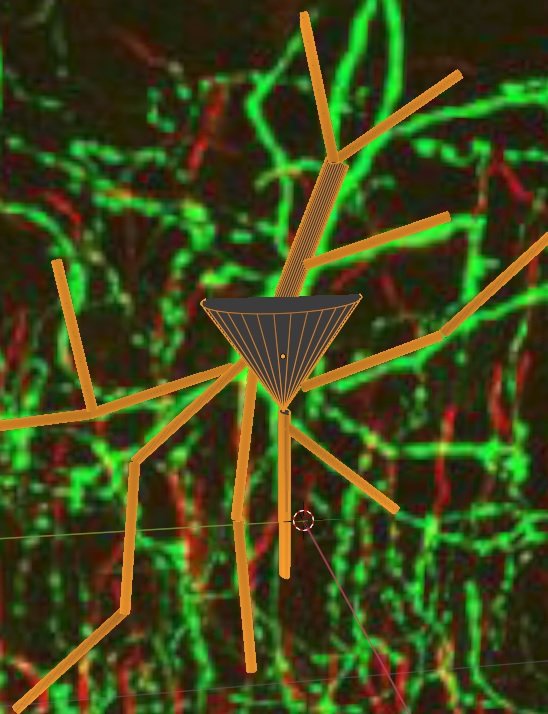
These methods are powerful, you can combine them with connection maps to achieve realistic brain geometry with very little effort. For example, Annie can "complete" a partially traced neuron, according to a branching spec. We haven't yet looked at what a branching spec actually "is", other than the locations of points, but Annie's branching specs are probabilistic and contain function modifiers. For example you can build this same chandelier cell using a "force of gravity" that directs the branches downwards, or you can apply any other function you can dream up. At the end of the day Annie understands these functions in terms of gradients, they're no different than a gradient of ephrins. As soon as I can keep VTK from crashing every half hour I can show you the end-to-end for how to do this in 5 minutes. Most of the time this extreme level of realism is not needed, it results in computational complexity with little added benefit. Lego-type neurons are often quite sufficient, they can be generated in seconds rather than minutes. Here's the same neuron, exported as an STL file and displayed in Microsoft's 3-D Viewer.

Apparently, Blender didn't do a perfect job of healing the mesh, but it doesn't matter. On the way back in, when this is imported into Annie, the faces will be reconstructed and the missing edges recovered. Here's the before and after. After Blender's "join" operation, the segments are joined but not necessarily in the right way.

After importing the mesh into Annie, this is the result - Annie has extruded the required components and now she'll reposition the vertices just like Blender does, only she'll do a better job. Now... there are many other ways of doing this. For example, some neuroscientists use the Neurolucida 360 software to trace neurons from microscopy. But it just exports OBJ files, it won't give you a much better mesh than Blender will.

I'd like to show you one of Annie's simulations, so you can see how that part works. The simulator is only there to test the networks, but it's pretty powerful, it rivals many of the others in terms of capability. The best thing about it is it's seamlessly integrated with the geometry. You'll see this on the next page, where we'll set up some CONNECTIONs so we get our model network to generate some data. (So far we have an axon but we haven't connected it to anything yet - and we can't do much with an axon without synapses!) You can see how Annie's neurons work though, they're meshes defined by simple geometry, and you can put things inside them (microtubules, membrane proteins) by attaching them to mesh vertices, and you can connect them together using a simple definition language that lets you specify anatomical arrangements and fiber pathways. The process of building a network is reduced to a few mouse clicks, or a few lines in a definition file. "Give me 100 neurons, lay them out in a 2d grid, connect them to the 5 nearest neighbors in the target layer". Boom, done. Then if you want to get more specific, you can get as specific as you wish. You can define the location of individual mesh vertices if you wish. Annie will let you build a mesh from scratch, with VERTEX and FACE directives. But who wants to do that? Let Annie do the work, tell her what to do and go have a cup of coffee while she does the job. When you get back, your network will be ready and you can push the "Start Simulation" button.
Let's be perfectly clear, meshes aren't a panacea and they're computationally expensive. There is a downside to working with meshes, you get a bunch of big files and you need sophisticated visualization to look at them. So, only use them when you need them. Annie provides a seamless path from the simple to the realistic. Target whatever part of the spectrum works for you. Chances are, you'll raise the bar once you see Annie in action. Imagine for a moment, what Annie can do when mesh geometry is combined with AI analytics on microscopy. When you create a neuron from a picture you have to imagine what its 3-d geometry might actually look like, but when you create it from serial sections of electron micrographs there's no guesswork involved. Annie will map the coordinates into your network space exactly, and generate similar neurons for you, and place them in a structure you specify, and connect them the way you specify, ... There's very little Annie can't do. If you need a realistic simulation, this is the way to get it. Annie is the answer to the disparate ecosystem of computational tools and mesh tools that scientists have to use to get their work done. Annie will import anything from anywhere, and when the simulation is done she'll export it in any form you need. She'll take all those ion channels and map them into a very pretty heat map of your membrane potential across space and time. Visualize it any way you want, online, offline, in fact Annie has a workstation driven tick mode so the server will never outpace what you see on the screen. Annie has a world class feature set, it just needs to be made available to users via the dashboard.
(By the way, I'm working as fast as I can, I'm just one guy trying to capture all the wonderful things Annie can do, and I'm building this web site on a daily basis so please keep coming back for the latest greatest updates! And, I'm doing this on my own without any sponsorship or grant money, if you'd like to help you can donate to Annie in the store). Annie has existed in one form or another since 1993. She was originally written in C, to run on a Windows 7 machine. Then she went through an MPI incarnation to run on an HP Convex and a cluster. Now she's been fully ported over to Python and uses VTK for visualization, and at this moment a brand new user interface is being built using the Panel package from Holoviz, which so far is idiosyncratic but works acceptably well. If you're wondering about the difference in appearance between the menus and the VTK images, I'm experimenting with that right now. Panel looks a little childish compared to VTK, and there are other tools (like Trame) that provide a slightly more professional appearance, they're harder to use though (and they have more dependencies). The user interface issues are frankly considerably more annoying and time-consuming than anything inside Annie - Annie has already been field tested for 25 years, she lives in about a dozen laboratories all over the world, in various forms and customized in various ways. Annie was inspired by the Nova film "The Brain", which I worked on in the laboratory of Dr. Robert Livingston at UCSD. We used an Evans and Sutherland Picture System and a Talos digitizing tablet to trace serial sections of brain into a PDP-11/34. At the time the SGI company was right down the street in Sorrento Valley, they were developing high level graphic libraries for visualization - and today, Python is where you find these things, they're free and open source and it's mainly a matter of putting together a toolset that can help with the workflows. There is no one-size-fits-all tool, we have to use the tools that are available to us, and Annie addresses an essential gap in the toolsets, and simplifies workflows, and improves the rendering of results and the time-to-publication.
Graphics Is Fun, But Show Me Some Numbers |




















