Finite element analysis on nerve membranes is not a new idea. The compartmental models of Wilfrid Rall (late 50's and early 60's) are an early form of the mesh concept. Meshes are simply very small compartments, and there are things we can do with meshes that we can't do any other way. Meshes are reappearing in a big way in the world of neuroscience, as people realize that neural information is not just activity levels, and in fact the interaction between activity levels and the intra- and extra-cellular environments are what distinguishes neuroscience from machine learning. Finite element methods have had some significant successes to date, for example a recent paper describes how extracellular stimulation actually works on a neuron (Fellner et al 2022). Similarly, there is medical interest in the details of stimulation effects on peripheral nerves (Pelot et al 2018). The Pelot group at Duke University is a good read, they're doing a lot of work with these models, and you can find some of their software on GitHub. Finite element models are also one of the few ways to link mechanics and electrical activity (Vasas et al 2024). The concept of "simulating a nerve network" has already outpaced the current state of machine learning in many ways. Neuroscientists as a group are keenly aware there's a lot more to neural networks than just matrix multiplication. And people are starting to figure out how to model syncytia, where you have small synapses embedded into a much larger continuous network (Jaeger and Tveito 2026). Meshes and finite element methods are required for these kinds of investigations, as they extend well beyond the traditional view of neurons as point-like electrical generators.
So now on to the math. There is a fundamental difference between solving equations in neurons, and solving them on a mesh. They're only the same insofar as the simulator can crank through each point performing calculations. In neurons, there's only one membrane potential, but in compartments or on meshes, there's lots of them, and they all have to be calculated individually. How you approach this is a question of your workflow and your scientific need. Meshes take time. If all you want to do is a little machine learning with some geometry, you don't really need a mesh, you can just tell Annie to build a simple network of the kind we saw on the About page, and start your simulation. Neurons and synapses compute more or less the same way (since their membranes are similar), but when one starts looking around at how the simulators actually handle this, one becomes horrified. Refractory periods are artificial, they don't happen because of the membrane properties, they happen because of a computational rule that says "remain dormant for 1 msec after firing". Synaptic behavior is determined by exponents that represent the cumulative activity of thousands of receptors, ligands, and who-knows-what-else. In a way, this is like artificially staging the timing within your network. And when done this way, it doesn't tell us very much when something succeeds. It's like, "look, it works!" - well yeah, that's because you programmed it that way. When you get 200 Hz oscillations it's because there's a time constant or a synaptic weight somewhere that's forcing the network into the dynamic. Maybe the astrocytes are sucking all the calcium out of your extracellular space, whereas you're off trying to explain the lack of dendritic spiking with synaptic inhibition. Accounting for all the factors that can affect ion concentrations in a neural network is still difficult, and we need the tools to be in place when the science is ready, because biologists (with very few exceptions) are not computer programmers and they don't have a whole lot of time to spend learning about software. It's more like "can this tool do what I need", and what's needed these days is a lot of visualization and some easy-to-access domain-specific analytics.
The benefit of simple neurons is they're easy to calculate. The complexity of your network can be measured as the ratio of computer time to real time. If a computer tick represents 0.1 msec and it takes an hour to complete, you have a complex simulation. A few years ago, IAF neurons were state of the art, and there's not a whole lot to them, typically their equation is something like:
dV = 1/τ * (Vr - Vm)
so when your membrane potential Vm is above the resting potential Vr, you get a negative correction that forces it back down towards the resting potential. The time constant τ determines how fast the correction occurs, with a long time constant the incremental corrections are slow and it takes a while to get back to the resting potential. This is simple enough, and it's also where the problems start. Inputs are currents, they're movements of ions from one side of the nerve membrane to the other. The movements determine the concentrations on each side of the membrane, and that's what results in a trans-membrane electric potential (a "voltage"). A "voltage" represents a ratio of ions. If we add or remove ions on either side of the membrane, we change the voltage. This is a very different concept from directly linking synaptic inputs to dV. In some simulators, inputs are multiplied by synaptic weights and the result is directly applied to what would otherwise be a GHK equation:
dV = 1/τ * (Vr - Vm) + I(s)
where the I stands for synaptic input instead of ionic current - the assumption being that the synaptic input will directly drive the corresponding current. Which is not always a good assumption. If you have magnesium ions blocking your channels your synaptic behavior relative to the generated currents might be a little different than the equation. It's important to be able to match the equation to the behavior. There are always underlying assumptions, and one must be explicitly aware of them rather than trying to sweep them under the carpet. Even the computational method used to solve the differential equations can sometimes affect the outcome of an experiment, so it's important to have access to a large library of methods and to be able to quickly and easily access them. How do I tell Annie to change the computational method? Easy. METHOD EULER. That's it, that's all. You can use that at the level of the simulation, the network, the cell, the neuron, the synapse, the ion channel, and any other computational object you can attach to a mesh. RK4 is a popular method, because it offers a reasonable compromise between accuracy and performance. There are methods that are computationally precise and almost never blow up, but they can also slow down your simulation. So Annie offers you a choice, choose the method that best meets your needs.
Synapses are the same way. They're traditionally modeled as a biphasic (usually bi-exponential) rise and fall, which is reasonable as far as it goes but if you have modulators or other external factors influencing those times you'll need to account for them somehow. A typical synaptic profile might look like this:
S(t) = Smax * A [e -t/τ1 - e -t/τ2]
where t1 and t2 are the rise and fall times. It gets a little more artificial when one has to introduce synaptic delays, which usually end up like
e -(t-τd) / τr
As you can see, this approach involves a lot of assumptions and a lot of compromises. It works, insofar as it generates what look to be superficially realistic results. But they're not really good enough for comfort. They're good enough to look at and say "okay, it's possible", but they don't speak to how the biological system actually works.
A more refined approach to simulation involves biological realism. For example, in addition to traveling down the axon, action potentials travel antidromically ("backwards") up the dendritic tree and into the apical dendrites, where they affect the ongoing behavior of voltage-dependent calcium channels and so on. In the hippocampus this behavior guides the timing of action potentials relative to theta. The .1-msec travel time from soma to dendrite is significant, it changes the network behavior. On a typical pyramidal neuron there are tens of thousands of dendritic spines, some of which may notice the action potential - so the timing of the incoming volley is crucial. One can certainly engineer this, by programming the time constants and delays - and it is legitimate to ask about the range of time constants needed to support some desired behavior. But at some level programming synaptic delays is cheating, they should be self-determined, not programmed. And the only way to do that with any kind of precision, is using a mesh. Otherwise, we are assuming that each synapse is isolated, and each neuron is isolated, and that's almost never the case in real life.
In the interest of realism, when we look at a simulation we need to use all the same tools the physiologists use when they're looking at live brains. FFTs, power spectra, coherence maps, that kind of thing. Visualization. Sure, all these things are available in Python, if you're a programmer you can move files around and pretty much import anything into anything else. But it's a lot of work, and it takes time. Wouldn't it be great if all that stuff was in one place? And when you think about it, if you can organize a simulation this way, you can organize a live experiment this way too. And actually that should probably become an industry standard, one should never sacrifice an animal unless one has at least a simulation-level understanding of the experiment. A simulation should accompany every piece of biology, in today's world there's no excuse for anything less. I was just looking at a very nice piece out of Princeton, from the scientists who work on the the Drosophila simulation project (Sebastian Seung, and others). The combination of techniques they were able to bring to the table to accomplish their analysis is first class. They trained artificial neural networks to segment axons and dendrites from electron micrographs! Great stuff. Unfortunately they chose Brian2 for their simulation project, which is understandable because they somehow detailed 180,000 neurons individually and that's a lot of text even for a supercomputer. Annie's value-add is in the translation of geometry to functioning networks. It's abundantly clear by now that the human brain is modular, and equally clear that there are differences in the construction of the modules from one area to the next. The scaling is such that precise modeling has to cover about a dozen orders of magnitude, which is more than a float32 but less than the 128-bit IEEE format. One of the interesting and noteworthy facts about the state of the art in neural network modeling, is no one yet knows what the temporal resolution of a dendrite is. In theory, a group of neurons has a discriminatory capability of τr / N, where N is the number of neurons and τr is the refractory period. Can dendrites make use of this? No one knows. No one's ever done a simulation at that level. (It's an instant publication, if someone wants to - and it would certainly be an acid test for any simulator). People look at STDP with time constants in the millisecond range, but dendrites can use the same trick that owls use when they locate prey - they can use delay lines and overlapping Gaussians to achieve precision that's a thousand times greater than that of any single neuron. The instantaneous condition of a patch of nerve membrane is indeed very local. The action of the Schwann cells on Nodes of Ranvier is proof enough of this assertion. Astrocytes in the central nervous system are undoubtedly even more precise, regulating the local calcium milieu at the synaptic level and perhaps even beyond. There are hundreds of other interesting scenarios around spike times. Just the idea of plastic inhibition is good for several careers.
Look - this is Lava-DNF, one of many available neural network simulators that provide spiking neurons and connection geometry.

Looks just like Annie, right? Looks exactly like the picture on the About page. They're after the same thing - a heat map of network activity. And, you can see their grid, it's just like Annie's 2-D retina. Only, Annie has already scrapped this approach, because it's too... well... it looks like a toy. They're trying to use the same tools that Annie tried, with basically the same results. They work, but they smell funny. They look like they belong in a corporate boardroom or on Wall Street or something. Annie has moved on to VTK, because it's the only thing that will handle her meshes. These heat maps need to be smooth, otherwise we're limited to very small grids for visualization. And neurons in small populations have a lot of issues that have nothing to do with their ion channels, like edge effects due to connectivity. Unless these are carefully accounted for, they will affect the behavior of simulations. Large faces are problematic, the grid size needs to be very small, like those axon meshes we looked at. In the above rendering the size of the point cloud is maybe a few hundred points at most. Whereas Annie's axon mesh has 8,000 vertices, and it only represents a tiny patch of membrane. Each segment of axon in the Blender tracing has several hundred vertices, and that's only because I was too lazy to expand them. Annie has this wired. It's important to understand this in terms of mesh resolution. To begin with, Annie will give you the grid in an interactive form. You can look at it in cell coordinates, or line it up with network coordinates, or rotate it any way you want.
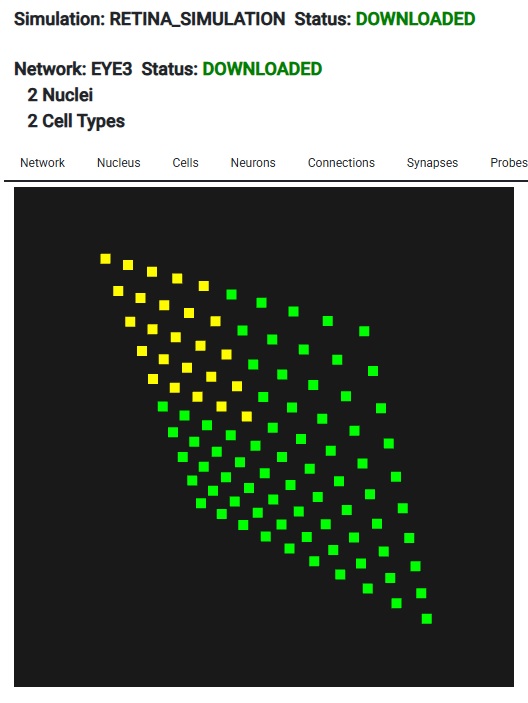 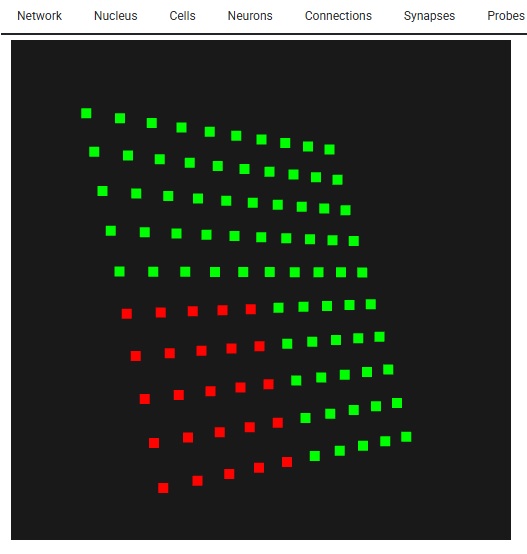
The important part of this, is we have a 10 x 10 grid of neurons - so when it comes to resolution, this particular form of simulation isn't going to do much better for us. If we had more neurons, we could do a little better in terms of mesh resolution, but as it is, if we asked Annie to create a very fine mesh for us it wouldn't do us much good. VTK becomes helpful with large meshes. For instance the WebGPU version of VTK can render several billion data points interactively on your PC screen, and in that case the mesh you'll see will be very detailed. Here's what 10,000 neurons looks like - it's not very many. With Lava we're SOL if we try to visualize these, however with Annie all we have to do is zoom in, and we get the exact same functionality, just with better resolution. The simple stuff matters!
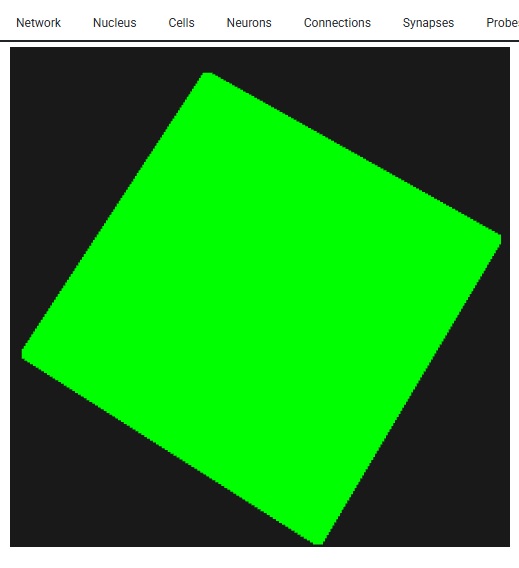
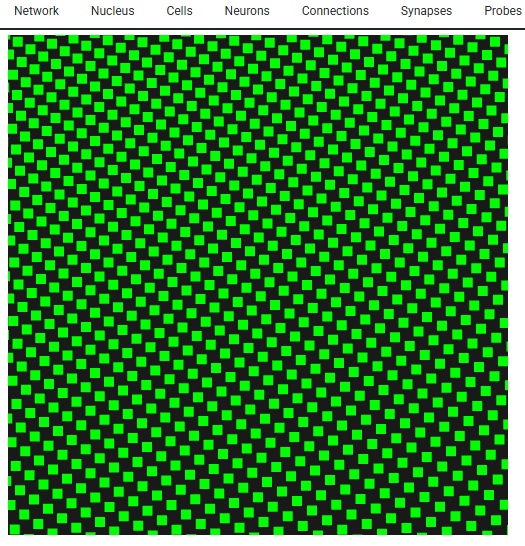
 
Anyway, hopefully you're getting a flavor for Annie and her approach to neural networks. Annie thinks like a neuroscientist, not like a machine learning engineer. Machine learning is a static exercise, whereas brains are real-time. The purpose of a brain is to optimize around "now". That kind of neural network, goes entirely beyond anything machine learning has to offer these days. Lately, we're beginning to understand some of the relationships between physics and the subjective world. But the basics still elude us. Why is red "red"? How come sensation is in my finger and not in my brain? Most of what simulations have told us so far relates to computation, like the information processing associated with the early visual system (which has certainly been one of the great successes of neuroscience). And currently, we can look at and even simulate the activity in the hippocampus related to memory and spatial navigation, but we can't say a word about the information content of the bursts between the hippocampus and the frontal cortex. Except to say that they matter. We'd like to be able to look at them, and understand how they convey information. To do that, we need visualization tools that go well beyond the simple presentation of anatomical voxels. We need meshes, and vertices with data attached to them, so we can accurately compute the information content of neural signals.
Some of the tools coming out of the E-Brains project in Europe are quite impressive, but they're all over the place, they're all different, they use different formats and data exchange is cumbersome. The big deal in the simulation world is visualization, the technology that does that will adopt itself, because people will use it, since it'll be the only game in town. There is nothing quite like Annie, she was a rock star in the supercomputer world when room-sized Convex FPS machines were popular, and I'm bringing her a modern dashboard because she can still do all the things people need, better and faster than her competitors. The mere idea of being able to set up 3-d network connectivity in seconds is a huge value-add, it can't be done this way anywhere else. How about a model of the corticostriatal connections that project into geometric clusters of spiny stellate cells? This geometry can't be achieved with "sheets" and "layers", but Annie can give it to you in seconds.
CELL NAME SPINY N_NEURONS 1000000 ARRANGEMENT CLUSTER_3D
CONNECTION FROM CORTEX TO SPINY TOPOGRAPHY POINT DIVERGENCE 100
Boom, done. Embellish as needed. Annie defaults in an excitatory connection type without plasticity, and topographically connects the cortical axons to the nearest 100 spiny cells (each). Now you have a 3-d topographical projection into a volume of neurons, which is something most simulators simply can't achieve. The simple stuff is important! Here's a cluster of stellate cells embedded into an array of projection neurons, arranged into "modules" of 1000 neurons each.
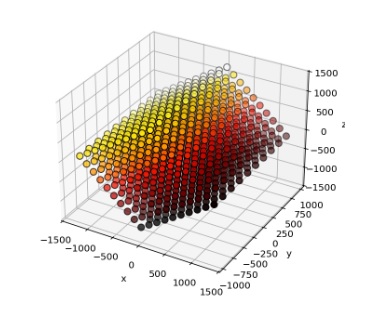 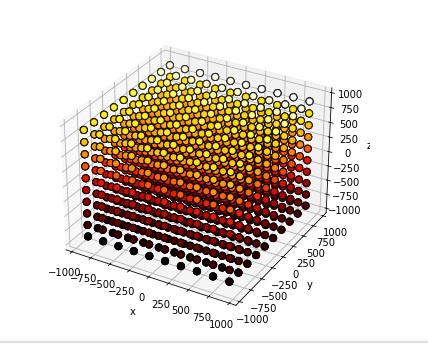 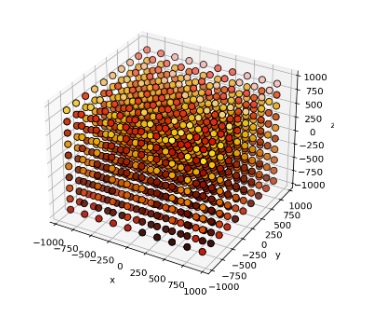
Time-to-build on this, on a Windows PC, is 0.59 seconds. That's 1 million neurons, with 100 million connections. This little gem chews up about 1.5 gB of computer memory. But it works, the first time and every time. And that provides a perfect segue into the world of developmental simulations, because they're huge and they take a lot of time. And they eat up a lot of computer memory. Often we find we have to swap things out, maybe keep some information in a database or something. That's how Annie acquired a database - a few years ago people were using her for developmental experiments and the memory requirements for thousands of growing meshes can get quite extreme, so we had to build some multi-processing capability to offload the intensive computations and add some "storage-swap" capability so meshes could be temporarily set aside while not in use. Annie's isn't quite as needful as Caltech's weather satellites but she can generate a LOT of output, very quickly. If you're running large simulations you should have at least 64 gB of computer memory and 1 tB of free hard drive. ("Large" in this case being defined as anything north of about 100,000 neurons - a fruit fly brain definitely qualifies). Give Annie the resources she needs, and she'll show you some awesome results. Annie is a seasoned veteran, she was already doing finite element analysis on meshed nerve membranes before today's young scientists were even born. One of the reasons she's alive is because Wilfrid Rall didn't have access to supercomputers. Right now, the direction is to integrate Annie's capabilities with AI. That marriage, will be very, very powerful.
This Is All Very Interesting, How About That Development Piece? |







