The issue with "all" neural network modeling is the user interface. Neural network modeling is a domain specific application, there is knowledge and there are methods that are in common use in workflows, and they're peculiar to neuroscience. If you're a doctor you look at a lot of MRI images and if you're a tax specialist you look at a lot of IRS rules, and if you're a neuroscientist you're interested in spike trains and extracellular potentials and sprouting axons and things other people don't know about and don't care about. The toolsets you need to use on a daily basis would mystify anyone else, even the simulation-heavy theoretical physicists will asks why you're doing things in peculiar ways, and the answer is because this is what the science requires. Visualization is absolutely the biggest deal in neuroscience. We need to visualize "everything", and my best advice after 40 years in the field is "don't listen to the naysayers". There is a famous story about Marvin Minsky and Frank Rosenblatt. As many of you know, Frank Rosenblatt invented the Perceptron in the 1950's, and he used simple neurons that were only slightly more advanced than McCulloch-Pitts neurons. He was working at Cornell and doing a lot of work for the air force, they were interested in analog computers for missile guidance and such - and Marvin Minsky was a computer scientist who looked at this from a raw computational perspective, and he wrote a paper explaining why Perceptrons can't do exlusive-or logic, and for some reason people listened to him and it set back the field back by 20 years. Meanwhile Rosenblatt had just created his first multi-layer Perceptron and shown it to the USAF and gotten a substantial continuing grant, and unfortunately he died in a boating accident shortly thereafter, so he was never able to counter Minsky's argument. However his scathing rebuttal does exist, it's buried in a classified DoD submission that was just recently declassified for public consumption. The reason I mention this, is because user interfaces for scientific modeling are much the same way. If you're a computer scientist trying to build a UI you get frustrated, because it's extraordinarily difficult. The tools created for the corporate boardroom are completely unsuited to a scientific need, and those created for scientific visualization are cumbersome and difficult to use. VTK is a beautiful rendering engine and it works on the web, but it's hard to use, you end up with 10,000 lines of code just to edit a mesh. Trying to put a VTK display inside a Panel dashboard doesn't work, the incongruity is simply too visible. Not to mention all the technical problems of re-rendering tabbed displays when reactive variables change. There's no way to use VTK to display forms, so when you're trying to build a dashboard you're stuck with merging two disparate technologies, one for the layout and another for the detailed visualization. This is one of the reasons why Annie is providing the first USEFUL user interface for neural network modeling. Think: you just built a model. Now you want to see a spatial map of the power spectrum. How do you do that? The old way was, you export your simulation results somehow, format them into a table (usually a file that can be read by some other software), read the file into Python or R, convert it to a dataframe, and plot it with matplotlib. That whole process can take DAYS, and that is a completely unacceptable situation in the modern world. If you're a scientist you need to be spending time in the lab, not in front of a computer terminal. The simulations have to be fast and the results have to be instantly accessible. Typically you'll want to start the simulation and return to the lab, and come back later when the computer's finished. At that point though, you'll want to see the results "right away". Otherwise you'll have to wait three months till you can hire an undergrad on work-study to do all the number crunching you don't have time to do. Here are some snapshots of a developmental scenario. It's hard to tell what's going on without an animation, and the user interface isn't fully ready yet so I can't show you the movie. (I could show you some VERY long tables that prove it's doing the right thing, but you probably don't want to see that). I'm working as fast as I can, this week I have an identical user interface in Trame and Panel, but neither one of them works. There's some silly stuff going on with the widgets, the tabs don't work right in either environment, and frankly I don't have time to become a widget expert, nor am I interested in debugging other peoples' tools. So my choices are, jump through hoops, or write my own display manager. (I've written display managers, they're not easy). The sad truth is that as of today, there is no way to quickly and conveniently build a scientific user interface. There is no standard, and no way to port between pseudo-standards. Annie aims to change that.  One of Annie's most important contributions is the idea of standardizing on a MESH as the fundamental unit of representation and computation. Rather than neurons, synapses, or artificially generated compartments. If you think about it, as a neuroscientist or a computer scientists, it's a natural. And it's one of the value-adds that distinguishes neuroscience from machine learning. A topographic neural network is best represented as a mesh, that way you can do useful geometry with it - otherwise why do topography, if you're not interested in manipulating geometry? The ecosystem of mesh tools is enormous, there is freeware like Blender that's every bit as good as the commercial products like Maya - and if you think about what these tools are actually doing, in terms of activities like rigging skeletons for gaming, a lot of this technology is directly applicable to neuroscience. We need to rig our cytoskeletons, don't we? They have limbs and joints just like real skeletons. And they move according to kinematic laws. Right now there is an explosion of technology in neuroscience, in the areas of recording and visualization, and the simulators need to keep up. Do these graphs look familiar?   We're in the world of large data these days, we can't run simulations on a Commodore-64 anymore. On the other hand, we don't need E&S Picture Systems either, because laptops and desktop PC's are very powerful and they can handle most of what we need to do - and if they're too small there's always the Cloud. The corporate boardroom-type visualization tools are going to stop working after 10,000 data points. They'll simply stop working - your task will hang up and that'll be the end of it, you'll have to kill it and start over. The workstations needs to be able to paginate through extremely large datasets consisting of millions of simulation points. A billion data points is not unheard of in a mesh simulation, the fluid dynamics folks and the rocket scientists have to deal with these volumes regularly. And the reality is, if your simulation is not at this level you're just playing around. Everything that comes out of neuroscience these days should be vetted by a sophisticated simulation, otherwise we end up with years of argument over issues that should be resolvable in seconds. Earlier I showed some simulation results from Donoso pertaining to hippocampal ripples, and there is still disagreement about whether the 200 Hz activity results from inhibition, excitation, or a combination of both. The answer is, it's not an either-or situation and the network behavior needs to be parameterized and quantified. For that you need a simulator. And I'm a big believer in the user interface, which directly determines the efficiency of the workflow in many cases. So let me just show you the kind of silly stuff I run into with these user interfaces. Here's one I did in Trame, looks pretty cool, right? It has the little navigation tray on the left, and some tabs, and a couple of dropdowns and some widgets on the right, and if you push the button you get a dialog box. Took me a couple of hours to throw this together from some examples. 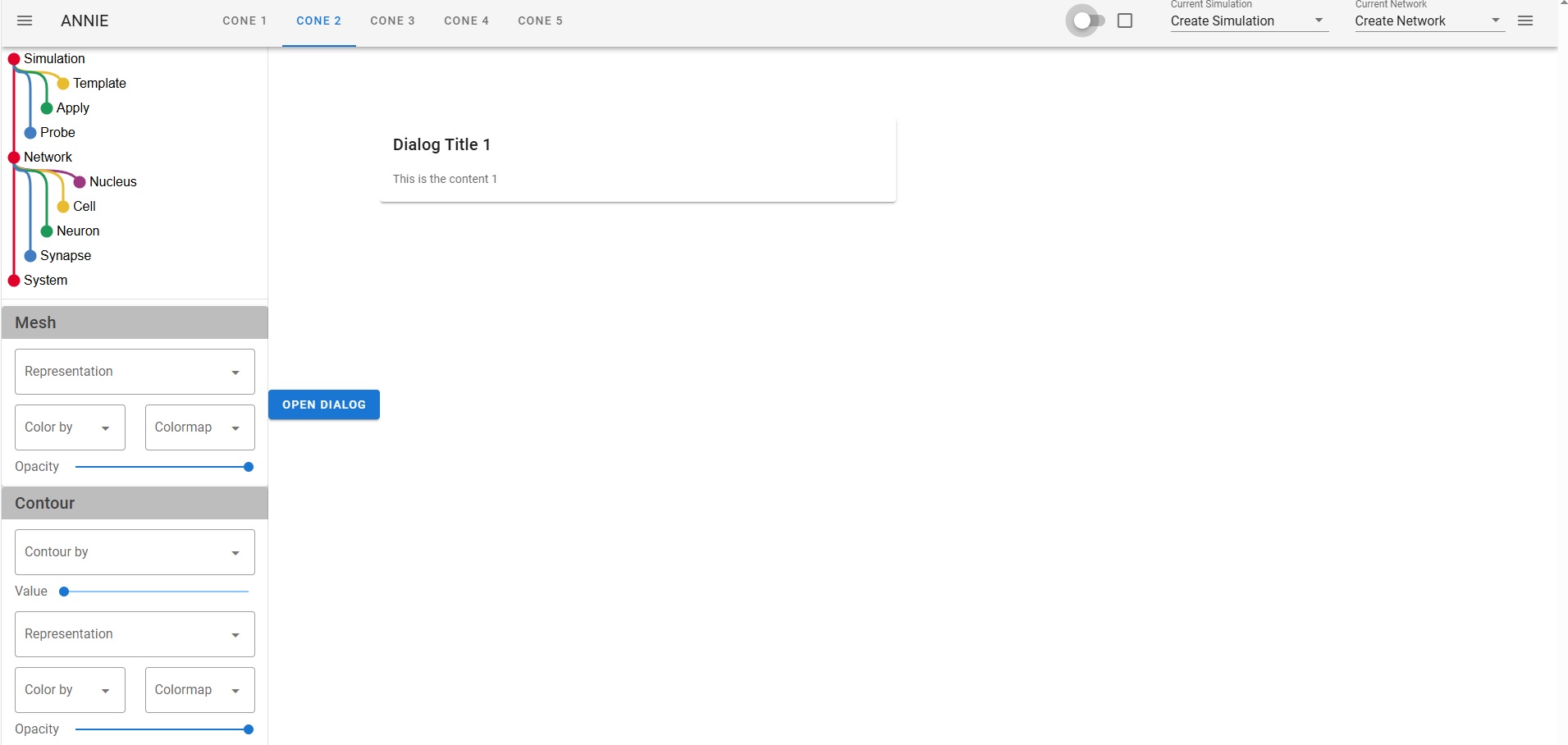 Look what happens when we make this dark, like we're going to display a mesh in color. Gee look, suddenly you can see the whole tab! But there are some problems here. First of all, those widgets on the right are supposed to be in the middle of the screen, just to the right of the tabs. The tabs are pushing them all the way over, and there's nothing I can do about it. If I put the tabs on the right they push everything over to the left. Over on the left, all the text in the navigation tree on the left sidebar is still in black, which makes it hard to read. The rest of the widgets change, but that one doesn't. So now if I want to make this usable I have to jump through all kinds of hoops to make the text white, but only when the user chooses a dark display. Or only when VTK is being displayed, or however it works. Silly stuff like this, is why there aren't thousands of excellent simulators out there. (Or at least it explains why people still use Annie in text mode, because she'll export a Pandas dataframe directly which is something most simulators still can't do). 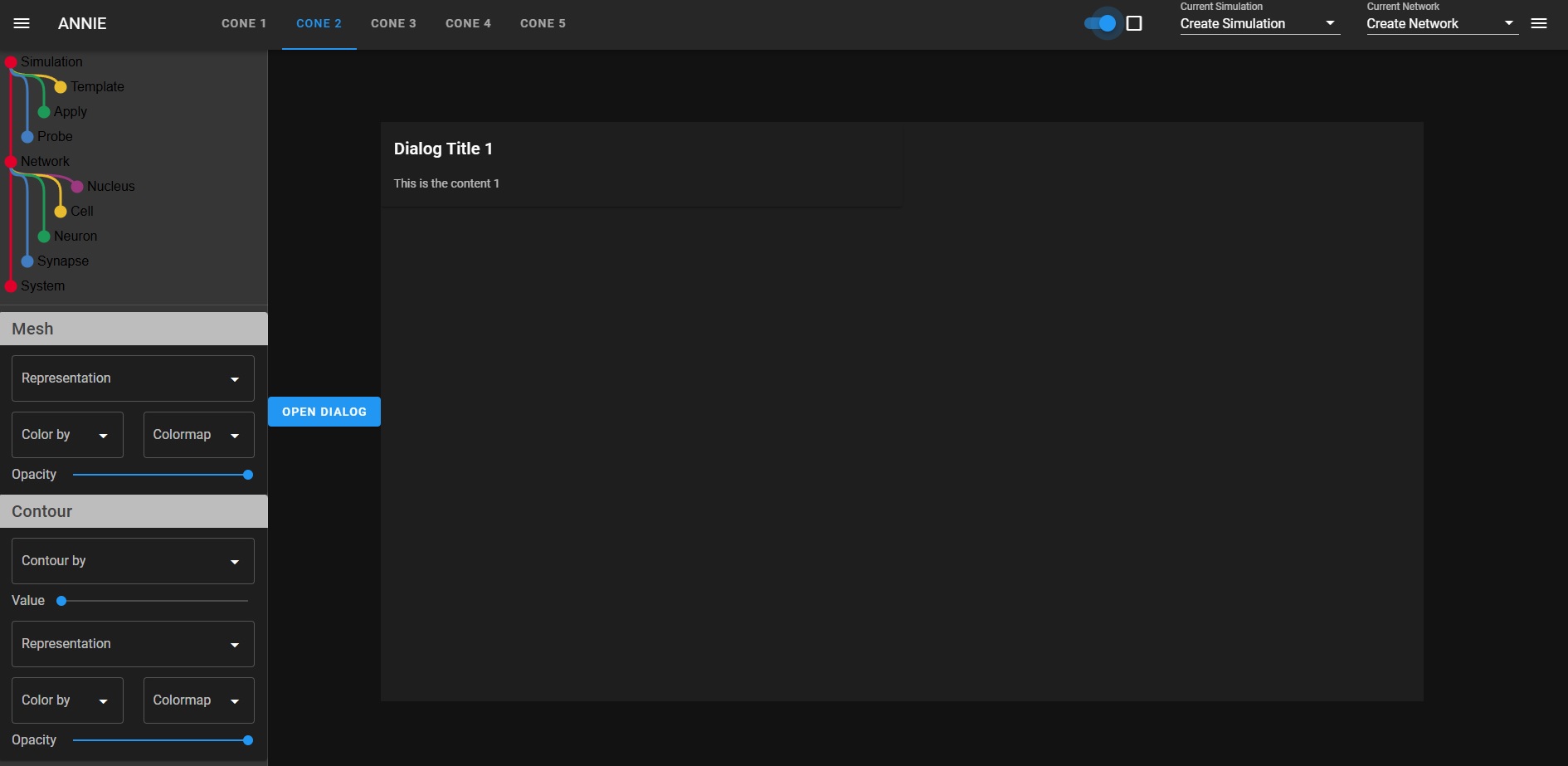 You can go look at some of the other "neural network simulators", many of them even explicitly say "we don't support Windows". The reality here, is that if you choose a user interface that can do justice to the visualizations, you're in application-land rather than on the web. It's do-able, in other words Adobe and AutoDesk and others have some wonderful applications that run on the web - but it took them years to accomplish and they're invested in specific graphics technologies. Trame uses something called Vue, which provides an enormous number of widgets but every time one of them changes, Trame has to change too - and if it doesn't keep up, the result is silly things like tabs that push other widgets over. And we definitely don't have time for stuff like this when we're trying to do science! Anyway, I'm going to stop complaining and let's do some science. Finite element methods are directly applicable to Hodgkin-Huxley axons. If you imagine a small patch of membrane that's defined by a piece of mesh, it may have a single ion channel in it, or perhaps two or three different kinds. That patch of membrane, has ion concentrations inside and out, and both the internal and external concentrations vary according to factors that have nothing to do with the membrane or its channels. Modeling the extracellular space is a non-trivial exercise. First of all cells are hairy, they have all kinds of molecules sticking up out of the membrane, and many of them are negatively charged which means the water molecules tend to align themselves in clusters around the charges, which in turn leads to further geometry, and all this affects the local ion concentrations by creating extracellular micro-currents. One of the purposes of glial cells is to control the variation of these micro-currents, and such wrappings can in fact be used for control purposes. All of this, is within Annie's domain, and none of it can be exposed with ordinary neural network simulators. The reality is the local membrane potential depends on ion concentrations, and if we're looking at tight geometry like in a dendritic spine, how are we going to account for the action of the glial membrane on the perisynaptic space? It's going to take more than synaptic weights to model that. People are building network models of the hippocampus without accounting for the behavior of astrocytes - which have gap junctions that form electrical syncytia and which transmit waves of calcium throughout the network. No one's looked at this yet, because the simulation is hard to set up. Not all the factors are understood - but that shouldn't prevent someone from beginning an investigation. Undergrads looking at a forest of divergent Python tools are going to slap their foreheads and get discouraged, but when they look at Annie's intuitive visualizations they'll say "hey, this is pretty cool, how do I...". Which is what we want, because no one knows how the astrocytes work yet. :) |