You can build networks three ways: interactively, with network definition files, and by importing networks from other simulators. Annie will support your workflow, however you like doing things. If you like working bottom-up, you can build entire networks interactively in the dashboard, even one neuron and one synapse at a time. When you're ready, push the "Build Network" button and Annie will automatically generate your network for you. Or, if you prefer working top-down, you can specify your entire network ahead of time, at any desired level of resolution.  |
You can see the example definition file here. This simple and highly non-biological "retina" is sufficient to demonstrate the dynamics of ganglion cell receptive fields. It generated the colored image on the About page, where you saw an interesting aliasing effect related to the hexagonal layout of the photoreceptors. Edge effects are important in small models, which is why scientists often prefer big models, they can ignore the edges and focus on the bigger middle. But edge effects become important (for example) when studying criticality, and in the retina in particular there is an interesting phenomenon during development where amacrine cells around the edges generate waves of activity that travel inward toward the fovea (thus mimicking "optic flow" long before the eyes open).
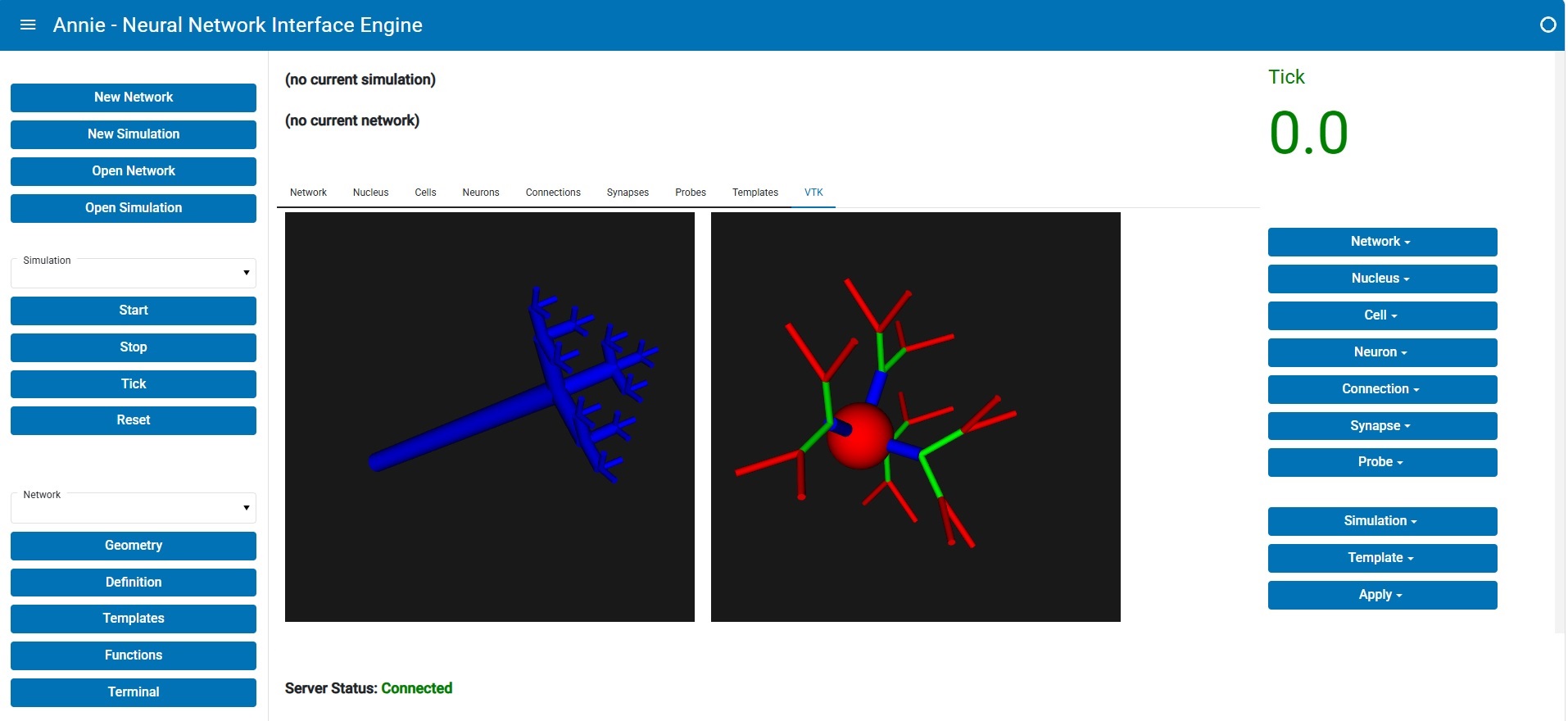
This is what Annie's client-server dashboard looks like after reading in the network definition file:
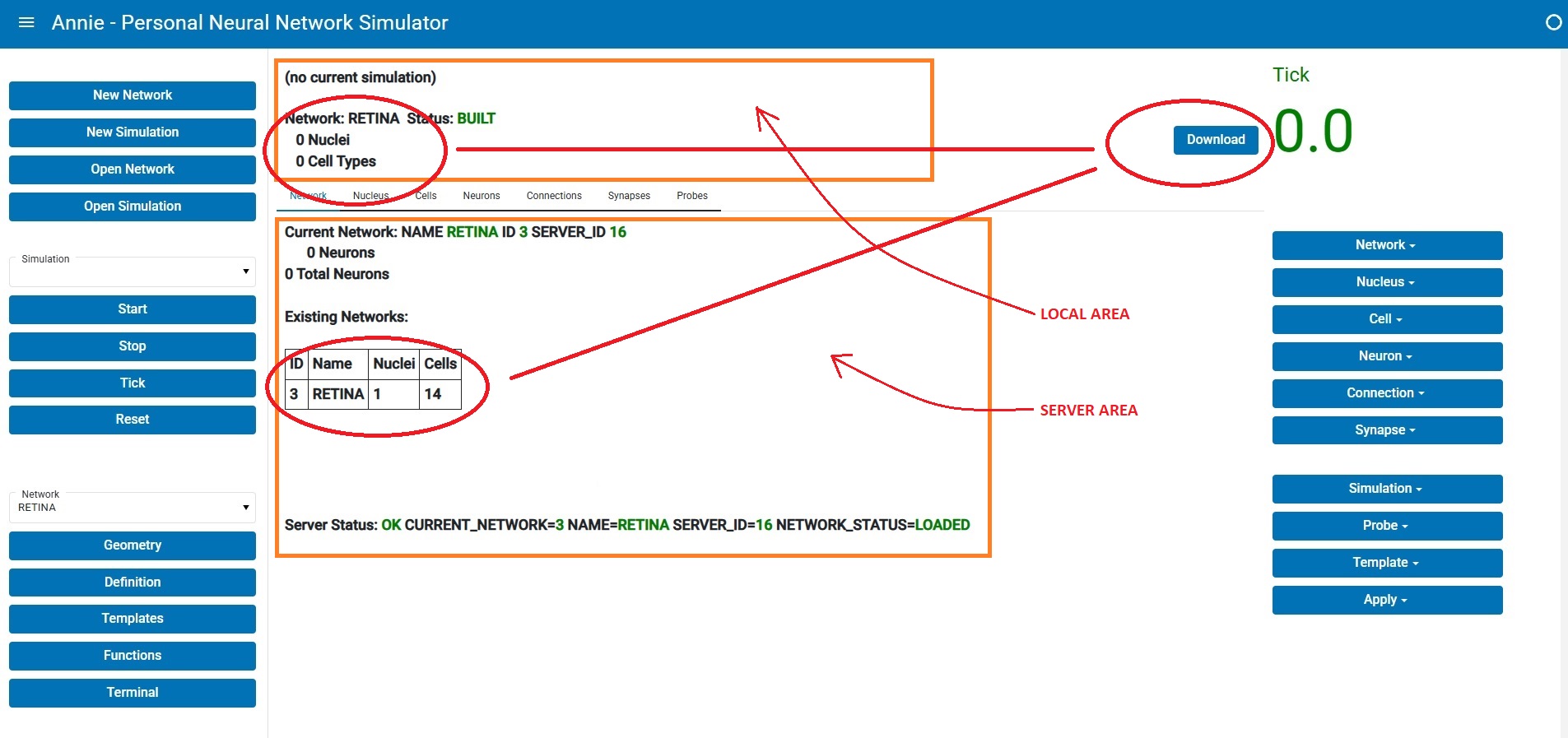
The middle pane is divided into two parts, a local part on top and a server portion in the lower half. You'll note that the numbers are different, and that's because the network hasn't been downloaded from the server yet. Annie is showing you a status that says "Built but not yet downloaded". (There are good reasons for separating the build and download functions). In the client-server configuration, the server builds everything and checks it first, before downloading the geometric information into the workstation. Once it's downloaded, the display changes and you see a quick report and a basic picture of what your network looks like. From here you can expand in any number of directions, and we'll look at some of them on subsequent pages.
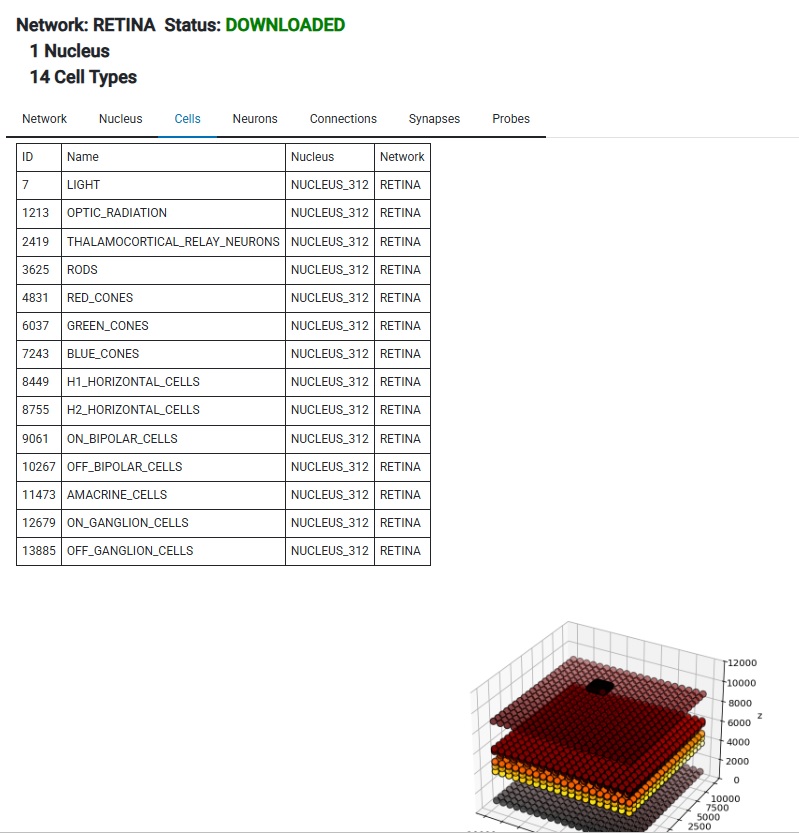
You'll note that Annie has created a nucleus for us, even though we didn't specify one in the network definition file. You don't have to specify a network or a nucleus in the definition file, everything is optional. This is a valid definition file, or it could even be completely empty:
CELL cell_name
We have declared a cell type without any neurons, and upon build Annie will dutifully build an empty network for us, giving us system generated names for the network and nuclei. This is a convenient way to generate a "shell" network that can be populated after the fact with cells of your choice. If you're running a series of simulations that require using the same basic network over and over again with slight modifications, this is a quick and convenient way to generate a folder full of experiments.
In the example network in the link above, we could have elaborated the LGN to include the interneurons, but we're just doing this for purposes of illustration. This file, while seemingly lengthy, is small compared to real (biological) networks. I didn't have to type it, Annie did it for me. She exports the network on demand, in human readable and editable form. And speaking of exports, this is one of Annie's export menus:
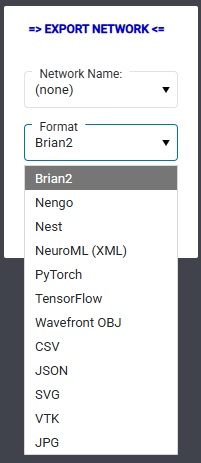
Hopefully you will notice and appreciate Annie's simple elegance. In the first release we've focused on the accuracy of reporting and the timely performance of simulations ("just the basics"), and in subsequent releases there will be plenty of embellishments - especially in the areas of graphics and visualization. Annie is a geometry tool, not fundamentally a simulator - even though the simulator has to be there to test the networks. If you're in the modeling business or you're serious about neural networks, you're doing it day in and day out, running dozens of simulations a day - you don't have time to scratch your head trying to figure out how to translate differential equations into idiosyncratic symbology. In many cases you'd be perfectly satisifed with an IAF neuron and a connection where you can just say "DIVERGENCE 5" and be done with it. Annie thinks like a neuroscientist, things are organized logically and geometrically based on workflow, because our modeling efforts rarely go from 0 to 60 in 3 seconds. Every one of Annie's directives is expandable, if you wish you can easily specify neurons and synapses down to the level of individual channel time constants and receptor subunit kinetics. But the present example illustrates a more basic approach to geometry, and quite frankly in many cases it's far more useful and productive than elaborating membrane properties. Some people have made entire careers out of studying receptive fields, and with Annie you don't have to sacrifice any animals to do it. Using Annie, you can build a full-blown early human visual system in about 20 minutes (if you know what you're doing), which is quite impressive considering there are about 180 million neurons involved. Annie is very efficient with computer memory, and for large simulations she offers both distributed multiprocessing capability and a database back end. If you don't mind waiting, you can run literally billions of neurons (not that it would do you much good, or let's say any more good than a simple million would).
So far so good? Everything looks vanilla, right? We can define a network, and it's probably good enough for some primitive computational purposes. But let's take it to the next level. We're doing finite element analysis on meshes, so we need Annie to create some meshes for us. Here are some simple meshes:
Annie uses primitives, like building blocks to construct more elaborate structures, and we'll look at how this is done on the next page. If we build a neuron this way, we may wish to lay out a collection of them geometrically, but we may also wish to connect them geometrically. Maybe the axons make a 90 degree turn after leaving their cell bodies, even while retaining their topographies. There are other ways of defining a retina, besides the simple linear example shown for the purpose of illustrating a network definition file. Annie will build a retina by replicating a "connection module", which is a small circuit you draw on the computer screen with a mouse (or you can type it too, but drawing is more fun). A module is a way of asking Annie to replicate the same structure hundreds or thousands of times. A cerebral cortex can be built the same way, the modules would be mini-columns, columns, and hyper-columns. In the superior colliculus the wide-field structure can be defined this way. You can define an entire network simply by defining the building blocks, and Annie will give you a fully meshed computational structure that resolves all the way down to the level of single ion channels. Meshes are powerful, we'll take a look at them in just a moment.
In the above example, we've laid out an overall geometry that puts light at Z coordinate 1000, and the optic radiation at Z coordinate 11,000. The inner plexiform layer is at z=4700 and the outer plexiform layer is at z=5800. The ganglion cells are at z=4000, and bipolar cells are at z=5000. Our photoreceptors are in the z=6000 neighborhood, and our LGN is around Z=9000. So we have established a basically linear pathway, from the light at 1000, to the optic radiation at 11,000. But, to be biologically consistent, we've placed the photoreceptors deeper than the ganglion cells, so light has to get all the way through the retina before it can impact the photoreceptors. And of course we've established a basic set of connections consistent with known retinal anatomy. This is a working network, we can actually run this, and when we do we can see the effects of the stimuli directly on the computer screen. We can ask Annie to build this network for us, and instead of simulating it we can request that its structure be exported in a dozen different formats. One of the useful things we can do with this, is bring the network into Maya or Blender, and convert all of the geometric structures to meshes, and then re-import them back into Annie. Annie will then dutifully interpret all of the geometry in mesh form, for example she'll calculate unions and intersections between meshes and so on. This would be a workflow for example, for an anatomist interested in creating a 3-d image of a brain structure. You can create a network inside Annie, export one of the cell types into Blender, changes its geometry, and import it back into Annie. Which means, you can trace the shape of your nucleus in the microscope, make a mesh out of it, and apply the mesh to Annie's cell group geometry. This is an enormously useful workflow with layered brain structures like the LGN and the superior colliculus. You can start by defining your LGN as a cube with EXTENT (10000,10000,10000), import it into Blender where it will show up as a bunch of modifiable vertices and faces, bring your microscope image into Blender and align it with your network object, and simply move the vertices until they match the microscope image. Then re-import back into Annie, and now you have a correctly shaped and scaled brain structure, and this is also a great way to properly define the outlines of layers and so on.
Here's an example. We can take the retina we just defined, and export it from Annie in Wavefront OBJ format, then import the OBJ file into Blender. When we do that, Blender shows us exactly the same objects as Annie does.
We can then make full use of the powerful mesh geometry inside Blender. Here is what our retina looks like, viewed from afar, complete with light source and a remote sheet of LGN neurons. Here we're displaying the network in wire frame mode, whereas the individual cell layers are shaded.
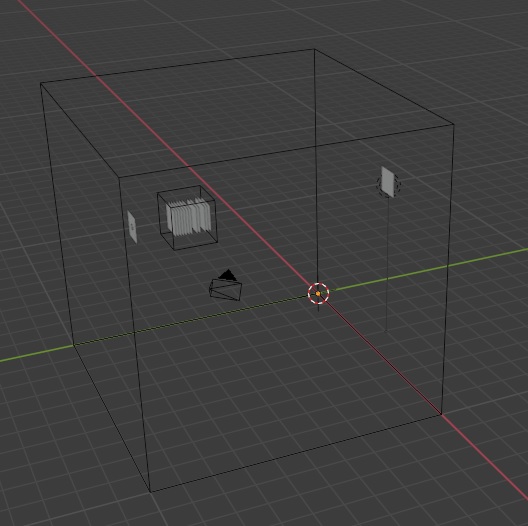
In this case, the effort is to manipulate the geometry. So we first tell Annie to create some cell layers for us, and without further instructions Annie will create them as rectangular grids (they show up as "cubes" inside Blender). Once in Blender, we can pick any vertex and move it, and the instant we do that the entire geometry turns into a mesh and Annie will read it back in that way, instead of trying to interpret the network in its original form. This is a powerful method for aligning models with brain maps - with some practice a 3d alignment can be accomplished in less than 5 minutes. You can make your network model "absolutely" anatomically accurate, to whatever degree you wish. Annie has a plethora of tools for arranging your neuron layouts to match meshes and all manner of regular and irregular geometries, so you can precisely and easily specify gradients of cell densities and such.
Anyway, we need to stay with the basics for a minute, before getting scientific. The first part of the example defines the cells, the second defines the connections. You'll notice we've made light inhibitory on the photoreceptors, and everything in the feed-forward pathway has a divergence of 1, whereas everything in the lateral pathways have higher divergences. Also the H1 horizontal cells and the amacrine cells have axons, whereas the rods and bipolar cells don't. A divergence of 5 means "connect to the 5 nearest neighbors" in the target cell group. Nearest neighbors are determined by registering the target coordinates with the source coordinates. Divergences can also be specified in terms of coordinate extents, like with the amacrine cells in the above example. A divergence of (2500,2500,5) means connect to anything within a radius of 2500 in the x-y plane, and we give the Z coordinate a little slop just in case we've declared the neuron locations with variance. One can also define a connection in terms of a MAP (as in the case of the on and off ganglion cells above). A map is like a function, or what other simulators sometimes call a "mask". A map is typically a function that defines how the connection should be made geometrically, for example a very common map is Gaussian connectivity, where the density of synaptic connections is higher in the middle of the target area, and falls off towards the periphery. A map is given by a .MAP file, and can be created and edited with Annie's map tool. One can also specify mapping functions directly using a FUNCTION directive, which is generally easier than creating a MAP. Maps are usually associated with complex axonal and dendritic branching patterns, and we'll look at an example on the next page, where we'll ask Annie to generate a fractal axon for us, and map it into a neuropil.
Annie can also generate connections dynamically. We'll talk about it later when we get to the network development piece. You can ask Annie to create constrained fractal branching trees for you, and program how they grow. One of the most powerful uses of this feature is in developmental models, where axons are sprouting and synapses are being pruned. Annie is completely aware of moving geometries, there is a large library of behaviors related to ephrins and etc. For the present purpose of simple illustration, you can create connections on the fly in Annie's dashboard, where you'll see a form like this:
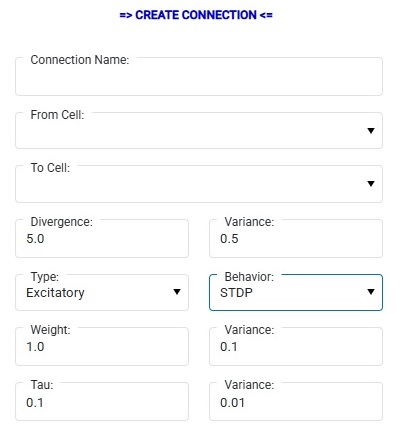
You can specify a connection this simple way, and there's a "+" button at the bottom if you'd like to engage in micro-management. A CONNECTION is from CELL to CELL, whereas a SYNAPSE is from NEURON to NEURON. There is also the concept of a "Projection", which is usually a fiber bundle from one nucleus to another (so a PROJECTION object is usually associated with a BUNDLE object, the latter being a geometric collection of axons that are constrained along certain paths). During the build, NEURONs inherit the properties of their parent CELLs, and synapses inherit the properties of their parent CONNECTIONs. SYNAPSEs are computational objects as well as geometric objects, whereas BUNDLEs are exclusively geometric (so far). When Annie builds a network, she only builds the geometric part. The computational portion is built later, when Annie builds the simulation. However after a network build, all the synapses are in place and the underlying computational infrastructure is completely defined. You can see this when you switch to the "Neurons" tab, where you might see a view like this:
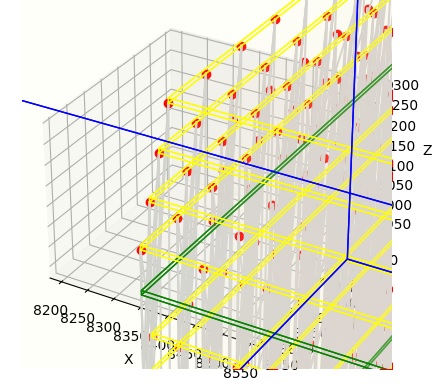
In this view we've only asked Annie to build synapses, without telling her how, other than by giving her the most elementary of CONNECTION specifications. Without further branching directives or a connection map, Annie will simply connect one neuron to another, and you can see the resulting axons in gray. This is an area where visualization immediately helps us, because one can tell at a glance the geometry is all wrong, there are fibers that travel right through neurons and all kinds of other non-biological horrors. However for computational purposes, this level of geometric non-sense might be okay, because we may not really care about the paths of the axons. However if we do care, we can ask Annie to fix these problems for us, by giving her slighly more refined CONNECTION directives. The idea is, you don't have to do more work than you need to. Annie likes simplicity, she's an excellent teaching tool for introducing young students to neural networks - because everything is interactive, with the dashboard you can change network parameters on the fly while simulations are running. In a more professional context you may only wish to do this in the early stages when you're exploring concepts, for publication a network definition file is more rigorous than sliders on a screenshot. But generally speaking, anything you can define in a network definition file, you can also do interactively in the dashboard. You can move seamlessly between one mode and the other.
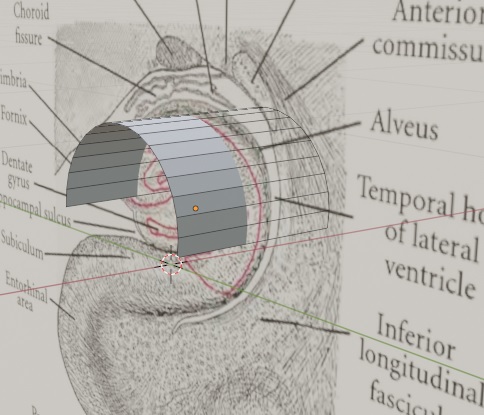
In the above example we have consistently specified TOPOGRAPHY POINT for the connections, this means they are point-to-point (topographic). This illustrates Annie's ability to line up geometries and perform calculations between them. There are many other ways to organize a network. You can have 6500 motor neurons in a clump of cells like the abducens nucleus, or you can have very sophisticated modular architecture like the cerebral cortex. Annie handles it all. You can get very specific with the geometry, you can define the shapes of cortical sulci and gyri if you wish, using splines or NURBs or meshes any other method the visualization tools will support. In the unlikely event that you can define them analytically by providing equations, Annie will handle that too. This is one of the beautiful things about meshes. You don't have to worry about equations describing your surfaces, you can simply draw them. If you have equations, you can use them, but the mesh doesn't require it. And meshes can be easily healed, if you're working with cylindrical primitives under a microscope you can join all the individual cylinders together with a single keystroke. We'll see an example on the next page. Meanwhile, here's a great example of the helpfulness of meshes, here I've imported a picture of a hippocampus, and arranged a curved plane in approximately the shape of the CA3 region I'm interested in modeling. It was easy, just create a cylinder, rotate it by 90 degrees, and move it to wherever the image says it should be - then delete the bottom half and the two caps. Less than 30 seconds from start to finish. Now I'd like Annie to populate "this" geometry with 100,000 pyramidal cells, and connect them topographically into the subiculum. At this point, Annie leaves the others in the dust. No one else can do this, in 3 dimensions or even 2, but Annie can do it in about a tenth of a second. All she needs is another similar sheet that describes the target. So in less than a minute, you've created some realistic connection geometry, which you can then embellish with basket cells and granule cells or whatever else you'd like to include. Maybe you'd like to tweak the apical dendrites a bit and put some voltage dependent calcium channels there. Easy - select a channel by clicking on a vertex, make the modifications, and tell Annie to apply them to "all" channels - as is, with a function, with a map, with a gradient or with a computed mesh, and with or without a variance.
You can see how the user interface for these choices gets a little complicated, there's lots of "+" buttons you can expand. For instance you don't want to see half a dozen fields related to variance unless you need them, so when the time comes you just click on the "+" button and the fields appear. Meanwhile you can still create objects just by pushing the "OK" button. Another of the very nice features of meshes is you can define functional compartments by simply defining regions with your mouse. You can imagine doing this with a Rall cable model. The old way, you had to define the compartments up front, and then build a mesh from the cable geometry. With Annie you can take any of the Lego neurons and zoom in on the axon hillock, define an area with your mouse and tell Annie "this is the hillock" and "please put some ion channels here". You can tell her it's a functional compartment if you wish, but she already knows, because the mesh has already defined it that way. Every face of a mesh is a functional compartment. The resolution of a mesh is so good that you don't need anything more - in fact with Annie's meshes you're already light-years ahead of the game, relative to Rall. Annie's mesh resolution is in the Angstrom range, she can visualize individual ion channels in a lipid bilayer. And if you need her to compute at that level, she can. This is why Annie has a server, because you may only be interested in a small patch of membrane, whereas Annie will generate enormous meshes, so big they couldn't possibly fit into a workstation.
One of the big problems with simulations is "too much data"! The moment you get into the simulation business, you'll find yourself overwhelmed with data. A 1-second network simulation using IAF (simple) neurons can generate over 1 gB of output when all the neurons and synapses are being read at each tick. So instead of reading "everything", we only read the things we're interested in, that way the web sockets don't get overwhelmed. To tell Annie you're interested in something, you can attach a "probe" to whatever you're interested in. You can attach a probe to just about anything. If you want a time series of the membrane potential of neuron 1641, you can attach a probe to that particular variable, and its value at each tick will be sent to wherever the probe says it should be written. It could be a file, it could be a web socket, it could even be a custom device in the operating system. If you'd like a time series of every neuron in a cell group, just attach a probe to the entire group, you can send the probe to a window to get a heat map or you can send it to a CSV file with columns that say "Tick" and "Neuron ID", and you can read that directly into Python as a Pandas dataframe and visualize it immediately using any of the popular plotting packages (Matplotlib, HVPlot, Altair, Vega, Plotly, etc). Or, you can just use Annie - she'll show you a 3-D rendering of your time series in an interactive window. When the server is running, you can scroll it around with your finger on a cell phone, on a PC you can use your mouse or the geometry widget. Annie has a Docker image too, she can be completely isolated from the rest of your computer and Docker will monitor things like memory and CPU usage while displays are rendering.